Our lab focuses on the development of new technologies for molecular analysis and biomedical research via advances in micro- and nano-scale sciences. We aim to develop new methods, devices and systems with unprecedented performance characteristics, such as sensitivity, specificity, resolution (temporal and/or spatial), multiplexing and throughput to rectify current technological limitations in the molecular study of diseases. In addition, we are moving beyond basic research into translational studies by developing and applying new technologies that address practical biomedical problems and clinical needs. By forging long-term collaborations with medical scientists and physicians across various disciplines (e.g. Oncology, Pathology, Surgery and Emergency Medicine) and by leveraging our engineering innovations in microfluidics, single molecule spectroscopy and functional nanoparticles, we are developing genetic and epigenetic biomarker-based diagnostics for cancer and an array of other diseases.
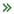 Quantum Dots for Genetic and Epigenetic Detection of Cancer
Quantum Dots for Genetic and Epigenetic Detection of Cancer
 Single Molecule Analysis of Genomic Content
Single Molecule Analysis of Genomic Content
Genomic analysis, such as sequence-specific nucleic acid detection and genetic mutation detection, has become one of the central themes for modern medical diagnostics and therapeutics. My lab has been developing new genomic analysis methods based on confocal fluorescence spectroscopy-enabled single molecule detection (SMD). We seek to take advantage of (i) the ultrahigh analytical sensitivity of SMD that allows amplification-free analysis of low-abundance nucleic acids and (ii) the digital nature of SMD that enables high-accuracy quantification of rare biomolecular targets. We designed a genetic analysis technology that detected genetic mutations according to two-color coincident emissions of single molecular probes that were resolved at the sub-millisecond resolution. This method expanded the capability of SMD for multiplexed DNA detection via combinational coincident analysis of multicolor fluorescent labels.
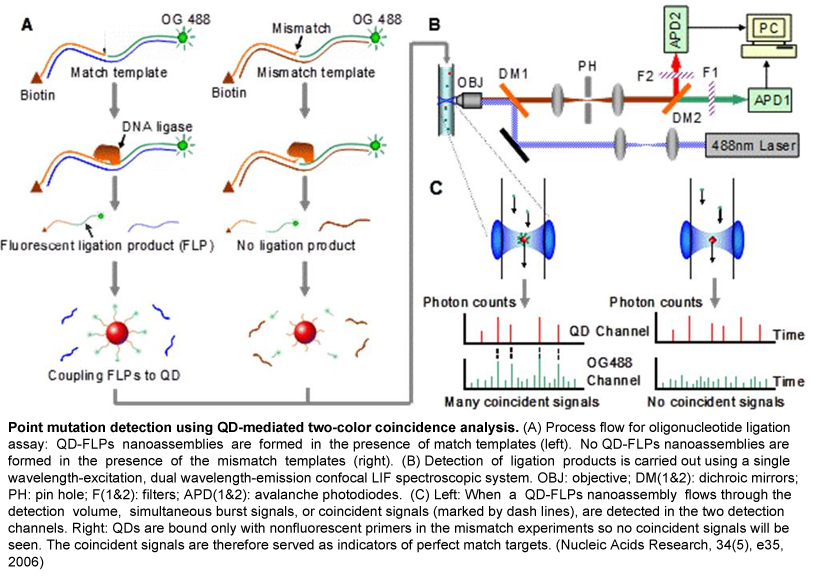
High sensitivity, molecular analysis using confocal SMD suffers from two prototypical technical limitations: low mass defection efficiency and quantification artifacts due to the minute, non-uniform Gaussian detection volume. We developed a new implementation of SMD, termed as cylindrical illumination confocal spectroscopy (CICS), which rectifies these shortcomings. As opposed to a diffraction-limited laser spot, CICS employs one-dimensional beam shaping to create a highly uniform sheet-like detection curtain that spans the entire cross-section of a microchannel. As a result, CICS achieves near 100% mass detection, is far less sensitive to thresholding artifacts than traditional SMD and is significantly more accurate at determining both burst rate and burst parameters.
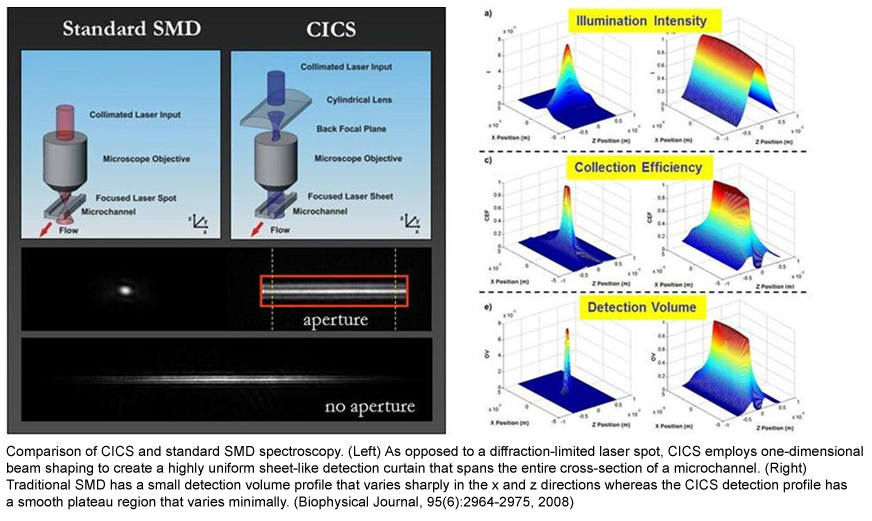
Using CICS, we developed the first PCR-free, single molecule assay to analyze circulating nucleic acids in serum and plasma. Circulating nucleic acids are a highly promising source for non-invasive biomarkers in a wide array of human diseases. The presence of long circulating DNA strands (i.e. increased DNA integrity) has been suggested to be indicative of neoplastic disease including cancer. Current PCR-based analysis of circulating DNA integrity is complicated by the limited sizing resolution, sampling error and shearing of DNA caused by DNA isolation. We developed a DNA sizing method that determines the size of a DNA strand according to the emitted photon count measured by CICS. This method can precisely and rapidly size and quantify DNA fragments directly in patient sera without DNA isolation or enzymatic amplification. This technique can potentially speed the clinical translation and validation of new circulating nucleic acid biomarkers since it offers researchers a simple, practical, and robust alternative to more complex methods.
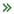 All-in-One Microfluidic Diagnostics: Sample Preparation, Target Detection and Integration
All-in-One Microfluidic Diagnostics: Sample Preparation, Target Detection and Integration
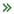 Droplet Microfluidics for High-Resolution and High-Throughput Screening
Droplet Microfluidics for High-Resolution and High-Throughput Screening

